Previous studies have established an association between gout and increased risk of cardiovascular disease (CVD).
Additionally, existing research has underscored the association between elevated serum uric acid levels and increased CVD and CV morbidity.
WHAT THIS STUDY ADDS
This study provides novel insights into the significant association between a high genetic risk for gout, distinct from hyperuricaemia, and an elevated incidence of CVD across diverse ancestries.
We further investigated the combined impact of genetic predisposition to gout and lifestyle factors on the risk of incident CVD using data from diverse populations.
HOW THIS STUDY MIGHT AFFECT RESEARCH, PRACTICE, OR POLICY
The findings of this study could lead to a deeper understanding of the genetic basis of gout in relation to CVD, emphasising the importance of polygenic risk scores in predicting disease susceptibility.
Clinically, these insights could guide personalised prevention strategies and inform public health policies to reduce the risk of CVD, particularly in populations with a high genetic predisposition to gout.
Introduction
Previous epidemiological studies have indicated that gout is associated with an increased risk of cardiovascular diseases (CVD) including coronary artery disease (CAD), myocardial infarction (MI), peripheral artery disease (PAD), ischaemic heart disease (IHD), congestive heart failure (CHF) and cardiovascular (CV) mortality.1–3 Therefore, the recommendations for gout management suggest that every patient with gout should be screened for combined comorbidities, including CVD.4 5 Until now, there have been several studies about the genetic relationship between serum uric acid (SUA) and CVD. Recent genetic studies for SUA have shown that SUA levels are associated with various phenotypes including hypertension, hyperlipidaemia, CAD, MI and IHD.6–8 Additionally, Kleber et al highlighted that SUA levels are related to CV and sudden cardiac death.9 However, despite the active research on the relationship between SUA levels and CVD, the genetic relationship between gout and CVD has not been sufficiently studied. Clinically, hyperuricaemia and gout are different phenotypes. Although hyperuricaemia is a risk factor for gout, only a minority of patients with hyperuricaemia develop gout. Additionally, asymptomatic hyperuricaemia generally does not require medical treatment, whereas gout often necessitates lifelong management.
The genetic architectures of hyperuricaemia and gout are significantly different. A recently published genome-wide association study (GWAS) study showed that 13 single-nucleotide polymorphisms (SNPs) reached genome-wide significance between asymptomatic hyperuricaemia and clinical gout.10 Thus, the genetic association between gout and CVD should be elucidated separately from previous genetic studies on SUA and CVD. To assess the genetic risk of gout as an indicator of CVD, we employed an approach using polygenic risk score (PRS) generation based on SNP weights derived from an independent large-scale GWAS.
In addition to genetic predisposition, lifestyle habits, including alcohol consumption, smoking, diet and physical activity play important roles in the development of CVD. Metabolic health profile is another key element influencing CVD progression. Furthermore, the effects of these factors on diverse populations remain unclear.
We investigated the association of genetic risk of gout and lifestyle factors with the risk of CVD and examined the beneficial reduction in CVD risk provided by two modifiable factors, lifestyle habits and metabolic health profile, with two independent datasets derived from different ancestral backgrounds.
Patients and methodsStudy population
The UK Biobank (UKBB) is a large prospective observational cohort study that recruited >500 000 adults across 22 centres located throughout the UK. The full protocol of the UKBB study is publicly available and the study design and measurement methods have been described elsewhere.11 Participants aged 40–69 years were enrolled between 2006 and 2010 and followed up for subsequent health events.
The Health Examinees (HEXA) Study was conducted according to the Korean Genome and Epidemiology Study (KoGES). The HEXA was a large-scale prospective community-based cohort study that recruited 167 169 male and female community residents in Korea between 2004 and 2013.12 The participants were followed up until the age of 80, and the ethnicity of the participants was Korean. Participants’ sociodemographic characteristics, medical history, family medical history and obstetric history were collected through questionnaires administered by well-trained researchers. Participants were asked to attend the participating centre for a follow-up assessment between 2012 and 2016.
In both cohorts, participants were censored for the occurrence of incident CVD or at the end of the follow-up. The study diagram is illustrated in online supplemental figure S1.
Definition of gout and comorbidities
Gout was defined by self-report at enrolment or by appropriate International Classification of Diseases (ICD) codes extracted from primary care or hospital records. Detailed definitions of the ICD codes can be found in online supplemental method S1.
Definition of cardiovascular outcomes
For the analysis of incident CVD and its association with Gout-PRS, participants with congenital heart disease were excluded to eliminate the possible association between congenital heart disease and CV outcomes (online supplemental method S1 contains the relevant diagnosis codes).
Incident CVD was defined as a diagnosis after enrolment in participants without pre-existing CVD including CAD, MI, PAD, ischaemic stroke and CHF. The ICD codes for each CVD are presented in online supplemental method S1. For each new-onset CVD considered, participants with pre-existing diseases at enrolment were excluded from the analysis. For example, participants with pre-existing any CVD at enrolment were excluded from the analysis of new-onset CAD, ensuring that recurrent CAD was not erroneously counted as new-onset CAD.
Definitions of variables
During the enrolment process in the UKBB, participants provided information on their sociodemographic characteristics, health/medical history and lifestyle/environmental factors through a self-administered touchscreen questionnaire and in-person baseline interviews.
According to the American Heart Association, five factors are primarily considered to define lifestyle behaviours: current smoking, alcohol consumption, obesity, physical activity and eating habits.13 14 Collectively, lifestyle behaviours are categorised into three groups: unfavourable (0–1 healthy lifestyle factor), intermediate (2 healthy lifestyle factors) and favourable (≥ 3 healthy lifestyle factors).15 Detailed descriptions and definitions of the variables considered in lifestyle behaviours can be found in online supplemental methods S2-4.
Metabolic health status was identified according to the presence of the five components of metabolic syndrome (MetS), based on the criteria from the International Diabetes Federation consensus report.16 Metabolic health status was categorised into three groups: ideal (0–1 MetS factor), intermediate (2–3 MetS factors) and poor (≥ 4 MetS factors). Detailed descriptions and definitions of the variables considered in lifestyle behaviours can be found in online supplemental methods S2,S5.
Genotype data quality control and imputation
Genotyping and quality control (QC) procedures and imputation followed standard practices and were performed using a cohort genotyping platform pair. We then generated each ancestry-specific genetic PC using QCed genotype data. Further details are described in online supplemental method S6.
UK Biobank (validation set 1)
UKBB samples (V.3; March 2018) were genotyped for more than 800 000 SNPs using either the Affymetrix UK BiLEVE Axiom array or the Affymetrix UKBB Axiom array.11 After QC and imputation, 377 909 European (White-British) participants were deemed eligible for validation genetic analyses.
Korean Genome and Epidemiology Study (validation set 2)
KoGES consists of 58 693 samples genotyped using the Affymetrix customised Korea Biobank Array chip (KBA chip, Affymetrix, California, USA).17 After exclusion, 58 686 Korean participants from the HEXA cohort were eligible for validation analyses.
Gout-PRS
The Gout-PRS was generated based on the large-scale gout meta GWAS summary statistics (6544 cases and 437 989 controls) from the GUGC (Global Urate Genetics Consortium)18 and FinnGen Consortium (Data Freeze R9)19 using the Bayesian polygenic prediction method PRS-CS.20 Individual PRSs were computed from beta coefficients as the weighted sum of the risk alleles by applying PLINK V.1.90 with the –score command.21 Details of the PRS analysis and GWAS summary statistics are described in online supplemental method S7.
Statistical analysis
Demographic and clinical characteristics are presented as mean±SD or number (percentage). Continuous variables were compared using Student’s t-test or the Mann-Whitney U test, as appropriate. Categorical variables were compared using the χ2 test or Fisher’s exact test, as appropriate.
To evaluate the risk of gout according to Gout-PRS, we used a multivariate logistic regression model to evaluate the association of Gout-PRS with gout. We calculated the OR and 95% CI after adjusting for age, sex, the first 10 principal components (PCs) of ancestry derived from genetic information, and the genotyping array type in the multivariate logistic regression model.
In the primary analysis, to determine the association between Gout-PRS and new-onset CV outcomes, multivariable Cox regression analysis was performed with adjustment for a history of gout and other covariates, including age, sex, PCs and genotype array, and HR and 95% CIs were calculated. The PRSs for gout were used as continuous variables. When categorized into groups, they were defined as follows: low (<20%), intermediate (20%–80%) and high (>80%). Subsequently, we conducted joint association analyses to investigate the interplay among genetics, lifestyle and MetS status. In addition, we performed sensitivity analyses according to CVD subtypes.
All statistical tests were two sided, and p<0.05 was considered statistically significant. All statistical analyses were conducted using R Statistical Software (V.4.1.0; R Foundation for Statistical Computing, Vienna, Austria) and PLINK V.1.90.21
ResultsPopulation characteristics
In total, 224 689 participants of European descent with available lifestyle/MetS status data from the UKBB (validation set 1) were included after excluding those with congenital heart disease or pre-existing CVD at enrolment. The mean age of the participants was 57.0 years (SD, 8.0 years). A comparison of the participant characteristics in each genetic group is presented in table 1. Among the UKBB participants, 4077 (1.8%) had a history of gout and 12 509 (5.6%) were diagnosed with incident CVD during follow-up.
Table 1
Characteristics of participants in the UK Biobank
For the cross-ancestry validation analysis, 50 364 participants of Korean descent from the KoGES (validation set 2) were included. Table 2 compares the baseline features and prevalence of gout in the study population according to the genetic risk for gout. The mean age of participants was 53.7 years (SD, 8.0 years). Among the KoGES participants, 347 (0.7%) had a history of gout and 673 (1.3%) were diagnosed with incident CVD during follow-up. The study flow chart for exclusion criteria is illustrated in online supplemental figure S2.
Table 2
Characteristics of participants in the KoGES
PRS with gout and validation in the KoGES cohort
We used large meta-GWAS (GUGC+FinnGen r9) data to compute the PRS for gout and found the Gout-PRS to be robustly associated with prevalent gout in the UKBB. Online supplemental table S1 presents the ORs for prevalent gout associations in the PRS risk groups. In the best-fit model, individuals with a high PRS had a 4.14-fold higher risk of gout (95% CI 3.69 to 4.64; p <0.001) than those with a low PRS group.
We validated the PRS for gout in a cohort from the KoGES. We found a significant association between PRS and prevalent gout across ancestry in the KoGES cohort (online supplemental table S2). Individuals in a high PRS group had a higher risk of gout (adjusted OR 2.59; 95% CI 1.80 to 3.75; p <0.001) than those in a low PRS group.
Association of Gout-PRS with incident CVD and its subtypes
In the multivariable Cox regression model (model 1), we found the Gout-PRS to be significantly associated with incident CVD in the UKBB online supplemental table S3. The association between Gout-PRS and incident CVD was maintained, and individuals with a high PRS had a 1.10-fold higher risk of CVD (95% CI 1.04 to 1.17; p <0.001) than those with a low PRS group (online supplemental table S3). In addition, we confirmed a significant association between Gout-PRS and incident CVD among some CVD subtypes in the UKBB cohort. The adjusted HRs were 1.14 (95% CI 1.05 to 1.24; p=0.002) for CAD, 1.17 (95% CI 1.00 to 1.37; p=0.046) for MI and 1.29 (95% CI 1.10 to 1.50; p<0.001) for CHF (online supplemental table S4).
In the validation set 2 (KoGES), we found a significant association between PRS and incident CVD across ancestry (online supplemental table S5). Individuals in the top 20% PRS group had a higher risk of incident CVD (adjusted HR 1.31; 95% CI 1.04 to 1.65; p=0.024) than those in the bottom 20% PRS group. Subtype analysis could not be performed because the only overall criteria for CVD definition were available in the KoGES cohort.
Joint associations of Gout-PRS, lifestyle and MetS with incident CVD
To explore the effects of lifestyle and MetS factors on incident CVD risk according to the genetic risk for gout, we stratified lifestyle and MetS factors by PRS category. We observed an incremental association among increasing PRS, lifestyle and MetS factors and a higher risk of incident CVD. Individuals with the top 20% PRS who had an unfavourable lifestyle had the highest risk for incident CVD (adjusted HR 1.99; 95% CI 1.73 to 2.30; p <0.001) (figure 1A), followed by those with high PRS who had a poor MetS status (adjusted HR 2.16; 95% CI 1.94 to 2.42; p <0.001) (figure 1B). In both analyses, P for the trends was significant.
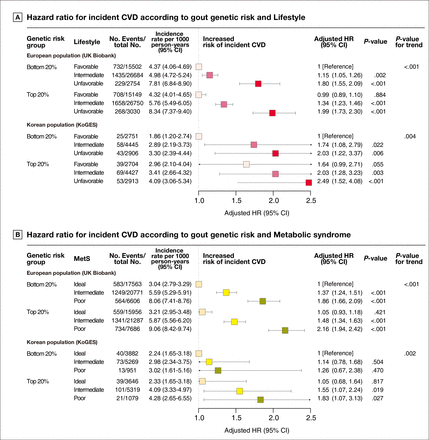 Figure 1
Figure 1
Forest plots for incident CVD according to gout genetic risk, lifestyle and metabolic syndrome. Cox regression analysis was performed with adjustment for a history of gout age, sex, 1–10 PCs, and genotype array. CVD, cardiovascular disease; MetS, metabolic syndrome.
For validation analysis, we performed a joint association analysis in the KoGES cohort. Individuals with the top 20% PRS who had an unfavourable lifestyle had the highest risk for incident CVD (adjusted HR 2.49; 95% CI 1.52 to 4.08; p <0.001) (figure 1A), followed by those with the top 20% PRS who had a poor MetS status (adjusted HR 1.83; 95% CI 1.07 to 3.13; p=0.027) (figure 1B). In both analyses, P for the trends was significant.
Impact of Gout-PRS, lifestyle and MetS on incident CVD
Participants with the highest expected risk those having a high (top 20%) PRS for gout, unfavourable lifestyle and poor metabolic health (MetS ≥4) exhibited substantially increased CVD risk, at 2.54 times higher than that of participants having the lowest expected risk (P for trend <0.001) (online supplemental table S6). Overall, adherence to a favourable lifestyle reduced the risk of incident CVD (figure 2). Subgroup analyses revealed that this risk reduction occurred regardless of the genetic risk group; that is, metabolic health and a favourable lifestyle reduced the subsequent risk of CVD by 62% among those with high genetic risk (top 20%) and by 46% among those with low genetic risk (bottom 20%) (figure 2). Even for participants in the low genetic risk group for gout who also had four or more components of MetS, adherence to a favourable lifestyle significantly reduced the risk of CVD by 25%.
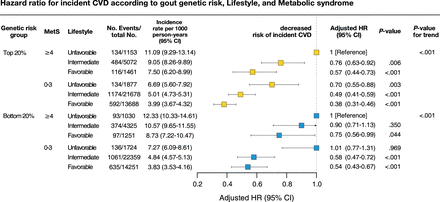 Figure 2
Figure 2
Forest plot for incident CVD risk reduced by lifestyle and metabolic syndrome in each genetic risk group in the UK Biobank. Cox regression analysis was performed with adjustment for a history of gout age, sex, 1–10 PCs and genotype array. CVD, cardiovascular disease; MetS, metabolic syndrome.
In the cross-ancestry validation analysis, due to the insufficient number of incident CVD cases, a joint association analysis using three factors was not possible, but the effects of lifestyle and MetS adherence on incident CVD were similarly identified, regardless of the genetic risk group. Adherence to a favourable lifestyle reduced the subsequent risk of CVD by 47% among those with high genetic risk (top 20%) and by 52% among those with low genetic risk (bottom 20%) (online supplemental table S7). Adherence to ideal metabolic health reduced the subsequent risk of CVD by 45% among those with high genetic risk (top 20%) and 17% in those with low genetic risk (bottom 20%), but the difference was not significant (p=0.569) (online supplemental table S8). Additionally, we investigated which factors could be more effective in reducing CVD risk by modifying them (figure 3). Notably, smoking cessation provided the greatest modifiable benefit in both multiancestry cohorts, reducing the risk by 41%–45%. Conversely, abstaining from alcohol was associated with an increased risk of CVD in both cohorts. The absence of a gout history resulted in a 30%–33% reduction in CVD risk across both cohorts. Furthermore, while most features did not show significant differences in impact on risk of incident CVD between the two cohorts, it is noteworthy that lifestyle modifications related to calorie dysmetabolism, such as ‘obesity’ management and ‘dietary (eating habits)’ control, and reducing ‘waist circumference,’ were more effective in reducing CVD risk in Europeans than in East Asians (as indicated by non-overlapping HRs and CIs) (online supplemental tables S9,S10). Lastly, we conducted sensitivity analyses to examine these effects within the top and bottom 20% genetic risk groups for gout. The findings demonstrated that the modifiable impact on incident CVD was generally similar across the genetic risk groups, and no significant interactions were identified between the genetic risk for gout and the various modifiable features.
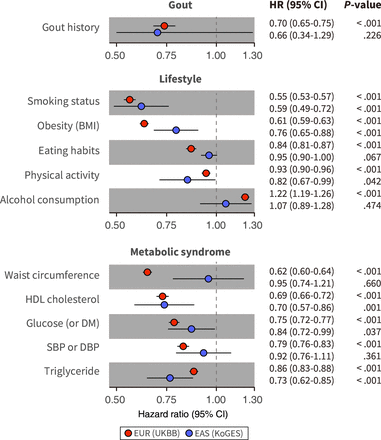 Figure 3
Figure 3
Impact of gout history, lifestyle and MetS factors on incident CVD across cohorts. Models were adjusted by age, sex, genotype array, baseline gout and 1-10 PCs. CVD, cardiovascular disease; MetS, metabolic syndrome; BMI, body mass index; DM, diabetes mellitus; SBP, systolic blood pressure; DBP, diastolic blood pressure; EUR, European; EAS, East Asian.
Discussion
In this study, we confirmed that a genetic predisposition to gout and lifestyle habits were significantly associated with CVD and validated the results in independent cross-ancestry cohorts. We showed that CV risk could be reduced through lifestyle changes, even in participants with a high genetic risk. Although there have been several studies on the relationship between the genetic risk for SUA and CVD, this is the first study to investigate the association between the genetic risk for gout and CVD. The risk of CVD increased with the PRS for gout, and this trend persisted even after adjusting for various covariates. The HRs for CVD increased further when lifestyle habits and metabolic health profiles were included, compared with when only the PRS for gout was used. This suggests a potential joint effect of these factors, emphasising their combined influence on CVD.
A previous study has revealed that the genetic risk for stroke and poor lifestyle habits increase the risk of incident stroke.22 Another study that analysed the data of 55 685 participants from three independent cohorts concluded that the PRS of CAD and lifestyle factors were independently associated with the risk of CAD. They also showed that a change to a favourable lifestyle can reduce the relative risk of CAD by 46% in participants with a high genetic risk.14 Unlike these studies that used a PRS for CVD, our study employed an RPS for gout, which predicted not only gout but also CVD incidence, suggesting shared genetic architecture between these conditions. Yun et al showed that the PRS for type 2 diabetes and lifestyle factors were associated with an increased risk of CVD.23 The diseases that increase the risk of CVD, such as gout and diabetes, may share genetic components with other CVD. Recent studies about PRS displayed that integrated PRS for multiple traits shows better predictive performance than PRS for single traits.24 25 The PRS for gout can be used as an additional screening component for individuals at high risk of CVD who cannot be found using the PRS for CVD.
Several mechanisms have been proposed to explain the increased risk of developing CVD with gout. Previous studies have reported that monosodium urate (MSU) crystals accumulate in tissues, including the heart and blood vessels.26 Dual-energy CT revealed MSU crystals in the coronary and aortic arteries.27 MSU crystal promotes oxidative stress and destabilisation of nitric oxide, leading to vasoconstriction and endothelial dysfunction. Inflammation augmented by uric acid, via the activation of inflammasomes or several inflammatory signalling pathways, contributes to the development of atherosclerosis.28 Intracellular accumulation of uric acid is believed to mediate oxidative stress in the mitochondria and has been associated with metabolic effects, such as insulin resistance, hepatic fat accumulation and the development of atherosclerosis.28 Besides the deleterious effects of uric acid, patients with gout usually have various comorbidities, including hypertension, diabetes, hyperlipidaemia, renal failure, urolithiasis and MetS. These combined disorders may contribute to the development and progression of CVD. Recently, metabolic dysfunction-associated fatty liver (MAFLD) has been reported as an independent risk factor for CVD.29 Elevated SUA contributes to a significant risk of MAFLD, and it shows a positive association with risk of CVD in patients with MAFLD.30 Therefore, gout and MAFLD might contribute to the development of CVD in a synergistic way.
Although epidemiological and in vitro studies have suggested that SUA levels play a role in CVD, the causal relationship remains controversial. An analysis of the Framingham Heart Study involving 6763 participants suggested that SUA levels were associated with an increased risk of CVD, but the association was no longer significant after correcting for other CV risk factors.31 In contrast, another cohort study that followed 5926 participants aged over 16 years concluded that the SUA level is an independent risk factor for CVD after correcting for other known CV risk factors.32 The results of genetic studies on the relationship between SUA levels and CVD are also inconclusive. A Mendelian randomisation (MR) study showed that HRs corresponding to each 1 mg/dL increase in genetically predicted SUA levels were significantly related to CV death and sudden cardiac death.9 However, another MR study reported that an increase of 1 SD in the SUA genetic score was not associated with diabetes, CAD, ischaemic stroke or CHF.33 On this issue, more evidence is required to reach a conclusion regarding this issue. Regardless of the debate over causality, it is clinically meaningful to use the PRS for gout to identify individuals at risk for CVD. If PRS is introduced into the practical field in addition to conventional risk factors, such as alcohol consumption, smoking, diet, lipid profile and combined diseases, it would be helpful to screen high-risk groups for CVD more effectively. To date, there have been several obstacles to applying the PRS in clinical practice. One of the barriers to applying the PRS to clinical practice is the lack of generalisability. Most PRS studies were conducted using a dataset of European ancestry, and the performance of the PRS was reduced in non-European populations. Therefore, it is necessary to develop a PRS for multiple ancestries to expand its generalisability. In this study, we confirmed that the PRS for gout is a predictor of CVD in Asian and European populations. Recent studies have focused on conducting PRS in larger and more ancestrally diverse populations. We believe that our results will contribute to expanding the generalisability of the application of PRS in clinical practice. PRS for gout can be used for preventing not only gout but also CVD. It is possible to identify individuals with high genetic risk for gout and strongly recommend modifying lifestyle habits. Weight reduction, smoking cessation, regular exercise and eating healthy food are effective strategies to prevent gout and CVD.
Our results showed that the change in metabolic health and favourable lifestyle reduced the subsequent risk of CVD by 62% in the population with high genetic risk (top 20%) and by 46% in those with low genetic risk (bottom 20%). The risk reduction effect of a healthy lifestyle on CVD was greater in the high genetic risk group than that in the low genetic risk group (62% vs 46%). Similar results were not observed in the validation set 2, which was thought to be due to the relatively small size of the validation set 2. However, the results of the validation set 2 also showed that a healthy lifestyle and metabolic health can reduce the risk of CVD in both the high-genetic-risk and low-genetic-risk groups. Our results suggest that intensive lifestyle modifications may be more beneficial in patients with a high genetic risk. These results are consistent with those of previous studies. Mega et al analysed the integrated data of 48 421 individuals from four cohort studies and showed that the effect of relative risk reduction for CAD by therapeutic intervention was greater in the high-genetic-risk group than that in the low-genetic-risk group (48% vs 13%).34 Another prospective study reported that the relative risk reduction of major adverse CV events by therapeutic intervention was greater in the high-PRS group than that in the low-PRS group (37% vs 13%, respectively).35 Summarising the above results, adherence to a healthy lifestyle was beneficial regardless of the genetic risk group; however, the benefits were greater for individuals with high genetic risk. Several recommendations about changing lifestyle habits can be suggested to reduce CV risk in individuals with high genetic risk of gout. Weight reduction is beneficial not only to reduce SUA levels but also to improve CV risk factors.36 37 The role of xanthine oxidase inhibitor (XOI) in preventing CVD is controversial. One cohort study with gout patients reported that XOI was not associated with an increased or decreased CV risk (HR 1.16; 95% CI 0.99 to 1.34) compared with those with untreated hyperuricaemia.38 However, another Japanese cohort showed that XOI was associated with reduced CV risk (HR 0.48; 95% CI 0.26 to 0.91) in patients with CV risk factors.39 The CVD prevention effect of statin in gout is not clear, but one study revealed that statin therapy has a protective effect in those receiving higher cumulative doses or with a longer therapy duration.40 Based on our results, smoking cessation, weight reduction, eating healthy food, regular exercise, control of blood pressure, glucose and hyperlipidaemia should be recommended for preventing CVD in individuals with high genetic risk of gout.
Limitations
This study had some limitations. First, this study showed the relationship between genetic predisposition to gout and lifestyle habits and the subsequent risk of CVD, but this does not indicate causality. Second, the definitions of lifestyle and MetS were different in each cohort because lifestyle habits, including dietary patterns, differ significantly between UKBB and KoGES. Therefore, it is necessary to develop a standardised method to evaluate lifestyle habits across countries. Additionally, lifestyle behaviours and metabolic health status were collected at enrolment, but these variables may change during the follow-up period. Such changes could potentially introduce bias into the results. Third, we did not provide direct evidence for improved outcomes of therapeutic interventions according to the PRS-based risk scoring system. A prospective study is needed to determine whether therapeutic interventions based on the PRS and lifestyle habits improve clinical outcomes. Finally, the limitation of our study was the use of the data, which differs significantly in ancestry and genetic architecture (UKBB and KoGES). While these differences pose challenges for direct replication, they also highlight the potential for developing and validating a cross-ancestry PRS that could be applicable across multiple ancestries across diverse populations. We acknowledge that further studies are needed to refine these methods and ensure that PRS can be effectively and equitably applied across different populations.
Conclusions
In conclusion, our findings establish a link between the PRS for gout, lifestyle habits and an increased risk of CVD using two separate databases representing diverse ancestral backgrounds. Additionally, we demonstrated that it is possible to reduce the risk of CVD by changing modifiable risk factors, lifestyle habits and metabolic health profiles. The effect of adherence to a healthy lifestyle was greater in individuals with a higher genetic risk. These findings suggest that therapeutic interventions based on the risk assessment of PRS for gout could enhance clinical outcomes through optimised personalised prevention.
Data availability statement
Data are available in a public, open access repository. All data relevant to the study are included in the article or uploaded as supplementary information. The PRS model constructed in the current paper is available for download from the GitHub page (https://github.com/dokyoonkimlab/gout-prs).
Ethics statementsPatient consent for publication
Ethics approval
All participants from the UK Biobank and Korean Genome and Epidemiology Study provided written informed consent, allowing the use of their samples and data for medical research purposes. This study adhered to the requirements of the Strengthening the Reporting of Observational Studies in Epidemiology (STROBE) statement.
Acknowledgments
This study was conducted with bioresources from the National Biobank of Korea, the Center for Disease Control and Prevention, Republic of Korea (NBK-2022-055). This work has been supported by the National Institute of General Medical Sciences (NIGMS) R01 GM138597. We thank the participants who contributed their data in the UK Biobank study. We want to acknowledge the participants and investigators of the KoGES.
Source link : http://www.bing.com/news/apiclick.aspx?ref=FexRss&aid=&tid=6706d7b43db0498fb2d6696343034802&url=https%3A%2F%2Frmdopen.bmj.com%2Fcontent%2F10%2F4%2Fe004552&c=839096489457485828&mkt=de-de
Author :
Publish date : 2024-10-03 17:00:00
Copyright for syndicated content belongs to the linked Source.









